Fit a Latent Dirichlet Allocation topic model using collapsed Gibbs sampling.
FitLdaModel(
dtm,
k,
iterations = NULL,
burnin = -1,
alpha = 0.1,
beta = 0.05,
optimize_alpha = FALSE,
calc_likelihood = FALSE,
calc_coherence = TRUE,
calc_r2 = FALSE,
...
)Arguments
- dtm
A document term matrix or term co-occurrence matrix of class dgCMatrix
- k
Integer number of topics
- iterations
Integer number of iterations for the Gibbs sampler to run. A future version may include automatic stopping criteria.
- burnin
Integer number of burnin iterations. If
burninis greater than -1, the resulting "phi" and "theta" matrices are an average over all iterations greater thanburnin.- alpha
Vector of length
kfor asymmetric or a number for symmetric. This is the prior for topics over documents- beta
Vector of length
ncol(dtm)for asymmetric or a number for symmetric. This is the prior for words over topics.- optimize_alpha
Logical. Do you want to optimize alpha every 10 Gibbs iterations? Defaults to
FALSE.- calc_likelihood
Do you want to calculate the likelihood every 10 Gibbs iterations? Useful for assessing convergence. Defaults to
FALSE.- calc_coherence
Do you want to calculate probabilistic coherence of topics after the model is trained? Defaults to
TRUE.- calc_r2
Do you want to calculate R-squared after the model is trained? Defaults to
FALSE.- ...
Other arguments to be passed to
TmParallelApply
Value
Returns an S3 object of class c("LDA", "TopicModel"). DESCRIBE MORE
Details
EXPLAIN IMPLEMENTATION DETAILS
Examples
# load some data
data(nih_sample_dtm)
# fit a model
set.seed(12345)
m <- FitLdaModel(dtm = nih_sample_dtm[1:20,], k = 5,
iterations = 200, burnin = 175)
str(m)
#> List of 7
#> $ phi : num [1:5, 1:5210] 5.77e-05 6.69e-05 5.73e-05 6.69e-05 5.28e-05 ...
#> ..- attr(*, "dimnames")=List of 2
#> .. ..$ : chr [1:5] "t_1" "t_2" "t_3" "t_4" ...
#> .. ..$ : chr [1:5210] "folding" "tosuprttedprtmnt" "importation" "hd" ...
#> $ theta : num [1:20, 1:5] 0.00043 0.23802 0.111462 0.210652 0.000615 ...
#> ..- attr(*, "dimnames")=List of 2
#> .. ..$ : chr [1:20] "8693991" "8693362" "8607498" "8697008" ...
#> .. ..$ : chr [1:5] "t_1" "t_2" "t_3" "t_4" ...
#> $ gamma : num [1:5, 1:5210] 0.188 0.19 0.207 0.206 0.209 ...
#> ..- attr(*, "dimnames")=List of 2
#> .. ..$ : chr [1:5] "t_1" "t_2" "t_3" "t_4" ...
#> .. ..$ : chr [1:5210] "folding" "tosuprttedprtmnt" "importation" "hd" ...
#> $ data :Formal class 'dgCMatrix' [package "Matrix"] with 6 slots
#> .. ..@ i : int [1:2864] 13 16 11 16 18 10 11 11 17 11 ...
#> .. ..@ p : int [1:5211] 0 0 0 0 1 1 2 2 2 2 ...
#> .. ..@ Dim : int [1:2] 20 5210
#> .. ..@ Dimnames:List of 2
#> .. .. ..$ : chr [1:20] "8693991" "8693362" "8607498" "8697008" ...
#> .. .. ..$ : chr [1:5210] "folding" "tosuprttedprtmnt" "importation" "hd" ...
#> .. ..@ x : num [1:2864] 1 1 1 1 1 1 1 1 1 1 ...
#> .. ..@ factors : list()
#> $ alpha : Named num [1:5] 0.1 0.1 0.1 0.1 0.1
#> ..- attr(*, "names")= chr [1:5] "t_1" "t_2" "t_3" "t_4" ...
#> $ beta : Named num [1:5210] 0.05 0.05 0.05 0.05 0.05 0.05 0.05 0.05 0.05 0.05 ...
#> ..- attr(*, "names")= chr [1:5210] "folding" "tosuprttedprtmnt" "importation" "hd" ...
#> $ coherence: Named num [1:5] 0.0817 0.395 0.24 0.0267 0.04
#> ..- attr(*, "names")= chr [1:5] "t_1" "t_2" "t_3" "t_4" ...
#> - attr(*, "class")= chr "lda_topic_model"
# predict on held-out documents using gibbs sampling "fold in"
p1 <- predict(m, nih_sample_dtm[21:100,], method = "gibbs",
iterations = 200, burnin = 175)
# predict on held-out documents using the dot product method
p2 <- predict(m, nih_sample_dtm[21:100,], method = "dot")
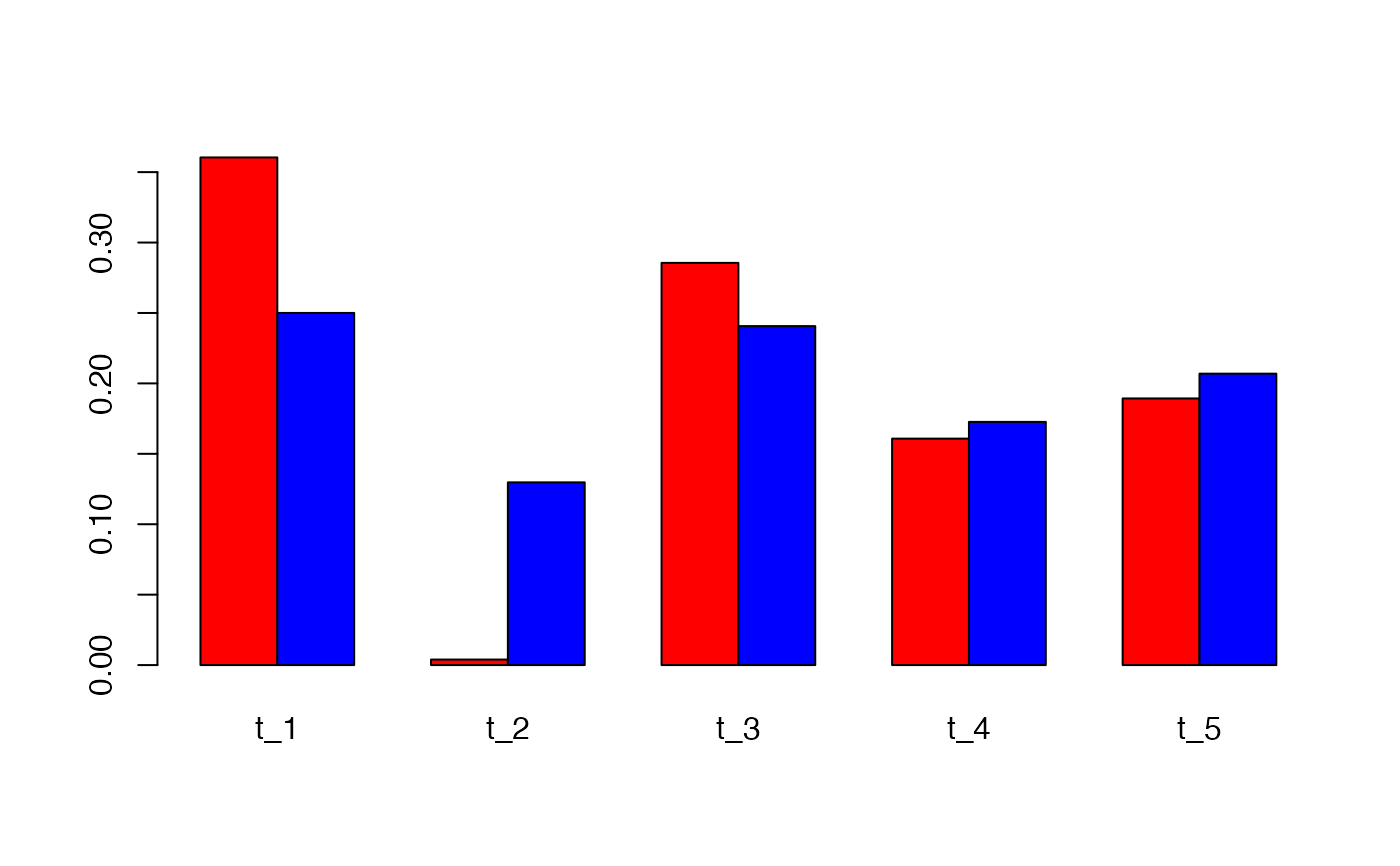
# compare the methods
barplot(rbind(p1[1,],p2[1,]), beside = TRUE, col = c("red", "blue"))